|
Neuroendocrine Prostate Cancer
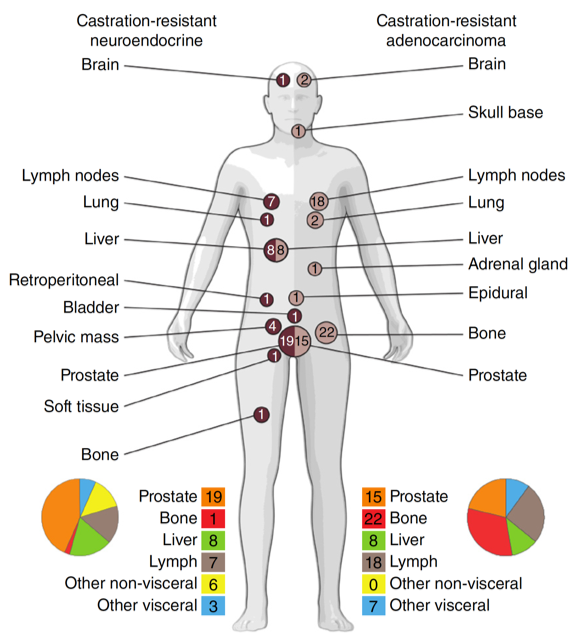
Lineage plasticity has emerged as an important mechanism of treatment resistance in prostate cancer, which may manifest as low prostate specific antigen (PSA) progression, resistance to androgen receptor (AR) pathway inhibitors, and sometimes small cell neuroendocrine pathologic features (Beltran H et al). At present, there is no established therapies for patients developing lineage plasticity or neuroendocrine prostate cancer (NEPC). Our laboratory is focused on understanding the biology underlying lineage plasticity that drives tumor evolution and therapy resistance. We have identified key genomic and epigenomic events that occur during NEPC progression (Beltran H et al), and we are exploring how the order of these events, tumor heterogeneity, and the pressures of AR-therapy influences plasticity and tumor aggressiveness. Our goal is to develop biomarker-driven clinical approaches for patients with NEPC through the initiation of early phase clinical trials. |
|
Organoid models
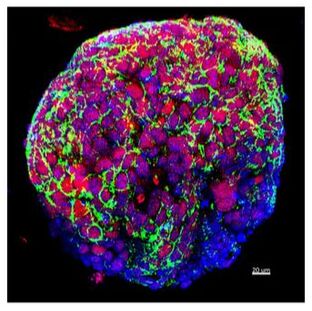
A major hurdle in the study of advanced prostate cancer and resistance mechanisms is a lack of existing preclinical models. Our lab has generated and characterized tumor organoids derived from needle biopsies of metastatic lesions. We have demonstrated concordance of molecular features between organoids and their corresponding patient tumors. These organoids also develop metastatic disease when transplanted into animals as xenografts. We are utilizing these organoids and organoid xenografts to understand the biologic role of epigenetic modifiers and other emerging drivers of NEPC progression and metastasis. High-throughput organoid drug screening nominated single agents and drug combinations suggesting repurposing opportunities (Puca L et al). |
|
Liquid Biopsies
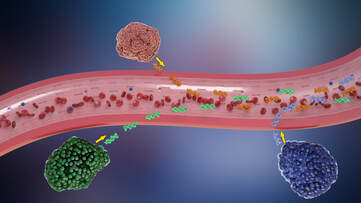
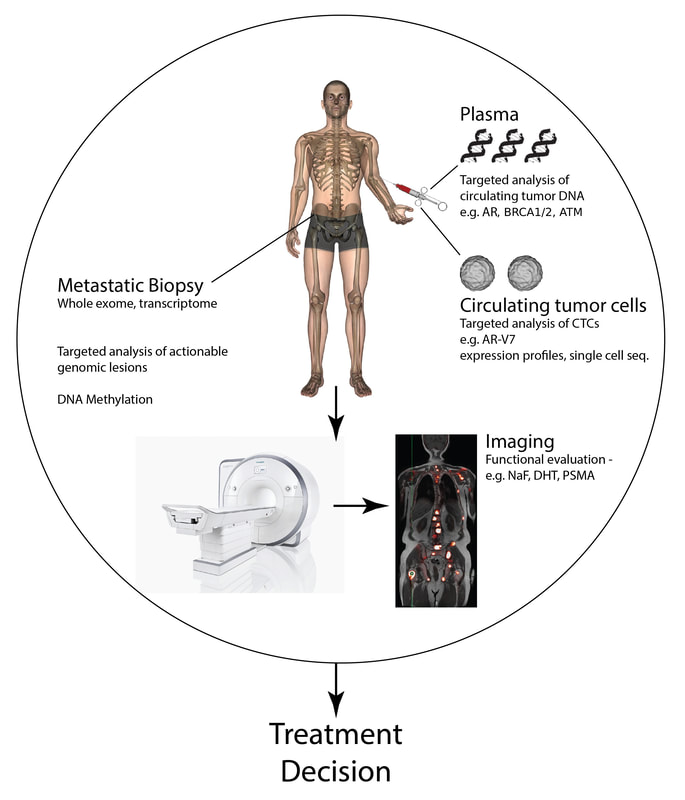
Serial biopsies to look for NEPC transformation or other mechanisms of treatment resistance are not always feasible or safe for patients with advanced prostate cancer. We are developing liquid biopsy approaches including circulating tumor cells and circulating tumor DNA (genomics and methylation) to detect the emergence of resistance features non-invasively. |
|
Precision Medicine Genomic profiling is widely used in cancer care to identify actionable alterations for individual patients within the context of precision medicine (Ku SY et al). However, not all patients with sequencing performed receive a targeted therapy, due in part to accessibility to clinical trials as well as a still incomplete understanding regarding the clinical, biologic and functional impact of identified alterations. We are working across cancer types to interrogate the genetic and molecular landscape of cancer, including the integration of newer platforms, such as whole genome sequencing, RNA-seq, and DNA methylation, to understand the functional and clinical significance of both common and uncommon molecular alterations and to develop new and informed therapeutic approaches for patients. Through collaboration within a multidisciplinary team at DFCI spanning both the clinic and the lab, we are working to develop the next generation of biomarkers and targets to inform cancer care. |
HOME | RESEARCH | LAB TEAM | PUBLICATIONS | NEWS | VIDEOS | SUPPORT US | CONTACT
©2022 Beltran Lab at Dana-Farber Cancer Institute
©2022 Beltran Lab at Dana-Farber Cancer Institute